bashML: Why Spark when you can Bash?
Technical
In one of our many research projects here at rev.ng, we are dealing with Big Data (is a 1..10 TB compressed database dump big? Well, probably not, but it is for us). Our first approach was to extract the data and store it in an SQL database, then run a bunch of queries and finally export the processed tables for other purposes. See the problem there? We used to use the database just like a, err... data processing tool? Unfortunately this wasn't working very well: we were having all kinds of performance bottlenecks since we were doing bulk inserts and bulk selects.
We then thought of using Spark or some other fancy stuff like that in order to stream process everything and just use text files. But, you know, we are a binary analysis company so most of the people here don't like garbage collectors (except me, the author of this blogpost, who like them very much). Anyway, we went from using MySQL to MongoDB+MySQL to MongoDB+PostgreSQL to, you guessed it, text files + good ol' Bash.
In this article I will persuade you, CEO at a brand-new Spark'ing startup that, sometimes, Bash'ing is all you need.
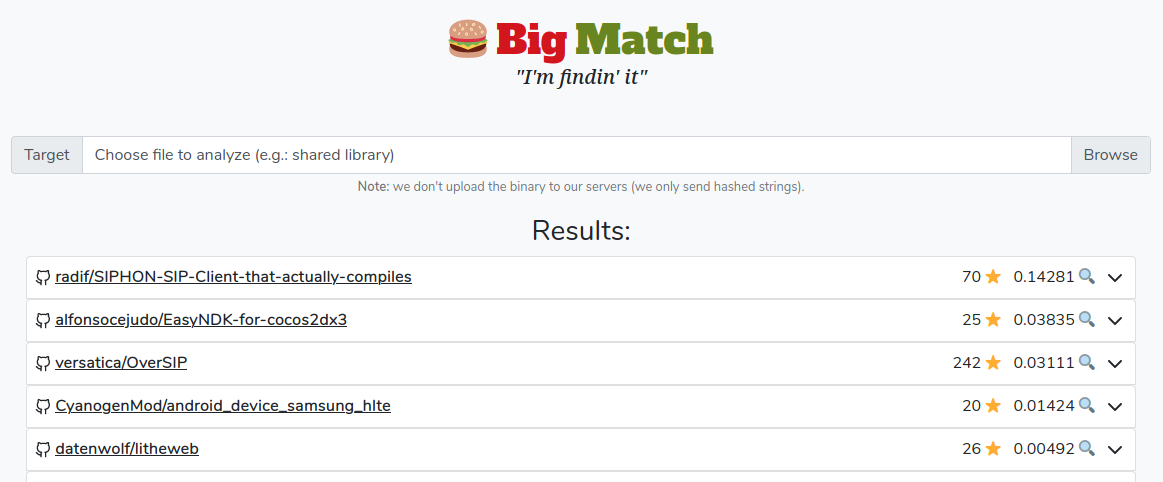
Note: If you haven't checked out our Big Match post, go read it.
An example dataset
For this tutorial we will be using a simplified version of our internal Big Match dataset of github repositories. It consists of a set of txt files and each of them contains a newline-separated list of sha256 hashes of strings found in a repo.
So for example, in file torvalds,linux.hashes.txt we have all the hashes of all the strings found in the linux kernel. They are also sorted for ease of use. It's clear that we use commas in place of '/' as a separator between username and repo name.
So, in this case, torvalds,linux.hashes.txt refers to the GitHub repo torvalds/linux.
Let's take a quick look at one of our files:
But before delving into the fun part, let's fix locale issues between tools. Trust me, you do want to do this, otherwise you will get all sorts of error when piping together different commands (e.g.: sort and join don't like each other very much).
Building blocks
Bash tools have all kinds of different options and there seem to be little to no consistency between them. We all know that: I know it, you know it, everybody knows it. So before starting we are going to create some helper functions for the most useful bashML operations out there.
We will start with a haskell-inspired helper, fst:
It's pretty simple: it takes the first column of a given file. By using $@, we can either give it a file argument or use it without arguments (think pipes) and it will Just Work™:
If you are wondering what bash does on <(...), look no further:
Long story short, bash replaces the argument with a file descriptor that's connected to the output of the command inside <(...).
In fact, you can also cat it:
With that out of the way, let's create some additional helpers:
For the few of you unfamiliar with tr, it's a little nice utility that can translate single characters (aka, map one character to another) or delete a character from a stream.
Another nice utility is nl, which counts the number of lines in a file.
Don't worry too much about the options: they are just there to output the format we want.
Example pipelines
It's now time to test our crazy bash skills to actually do something useful.
Repository similarity
Let's say we want to query our repos to find Linux forks just using common strings. We can use a scoring formula called Jaccard index (also called Jaccard similarity):
So, if and are our repositories, is a measure of how many strings they share, divided by the total number of their strings.
How can you do this in bash?
Using comm and some basic awk.
If you are unfamiliar with comm, it's a simple command that is able to compare sorted files line by line.
If you run it without any specific option it will output 3 columns formatted like this:
Remember to sort its input files otherwise comm will complain!
Anyway, if you run it with --total, it will also write a final line with the total count of lines for each column:
See that last line?
It's the count we were talking about.
We can throw away everything except that last line, do some simple math in awk, and get the Jaccard index of the input files:
Same thing, but with a different file:
In this examples, the first two files have only one line in common out of 5 unique strings, so . In the second example, they share 2 lines out of 4 unique strings, so .
Yes, this is what data scientists get paid to do by the way.
Now let's say we want to find the Jaccard similarity between Linux and the biggest repos in our dataset. We need to start by getting the list of such big files:
We now want to use the same jaccard() function we defined above but we want to run it faster by exploiting parallel processing.
We can do that with GNU parallel.
GNU parallel has a metric ton of options.
Yes, that is 1000-kilograms-many options.
Unless you voted for Brexit or play football with your hands: in that case it's ~2205 pounds.
Since we are Europeans and are not here to start a debate about our (superior) measurement system, we will just use 2 options and ignore the others:
What's happening here? Quite a bit actually: parallel just ran echo {} 3 times, every time substituting {} with a line from the input.
Remember: the calls to echo happen in parallel using multiple processes!
GNU parallel allows you to use {} multiple times if you want:
The options we give to parallel are very important: -j$(nproc) is for using as many processes as the processing units on the system (returned by the command nproc), while -k tells parallel to honor the input order when printing the output (so we don't get, e.g., c c c then b b b then a a a).
We can now use our previous jaccard function inside parallel to get the similarity between Linux and the other repos.
However, we also want a nice output so we throw in a call to printf.
The command gets a little messy due to string formatting, but just focus on the jaccard part:
We can see that repos that are just forks of the Linux kernel get almost a perfect score.
Then we have some repos that include the linux kernel but also have other stuff in them.
Finally, we have some random repos like kishikawakatsumi/Mozc-for-iOS that have nothing to do with linux.
With this information we can, e.g., do a simple similarity-based repository deduplication. In fact, at rev.ng we used this technique in one of our projects.
Repository vector encoding
What if we want to use some fancy deeplearning algorithms on our repos downstream? It would be convenient to have them represented as sparse vectors.
In other words, we want this
to turn into this
Where the numbers are just unique IDs for XXX, YYY and ZZZ.
The first thing we need to do is to get the complete set of hashes of strings in our repos. That's easy enough:
We then want to assign a unique number to each hash. This is how we can do it:
Let's generate the full (hash, hash_id) map:
We now need to apply this mapping to every file in the dataset.
For the sake of simplicity, we are going to focus on a single synthetic file called small/repo which we created by taking few string hashes from the linux kernel.
How do we use the mapping in hash-to-id.txt to get our vectorized version of the repo?
Enter the join command.
join is another nice Unix utility and it does exactly what it says: an SQL-like join.
Let's see an example:
This is your usual INNER JOIN, but in bash.
Using a similar command we can get our vectorized small/repo:
We now strip away the hashes to keep only the indices:
And there it is. It's trivial to load that up in a scipy sparse matrix, or similar, and do some proper machine learning using a proper garbage-collected language.
Bonus point: bash visualization
Ok, CEO at a brand new Spark'ing startup, you are ready to jump on the bashML bandwagon after what I've shown you, but you still feel like you are missing all of your web-based scheduling dashboards?
We've got you covered.
At rev.ng we developed a simple tool that uses strace to track the flow of data between different processes: pipedream.
It works by tracking file-related syscalls, along with pipes, and then trying to match I/O flows using file descriptors.
It's not perfect, but it works in most cases.
Let's put everything we've seen in the last part of this blogpost in a single shell file and use pipedream to get a nice pipeline visualization:
And this is the pipedream we get:
Some considerations
Before leaving you to your very much important CEO duties, I'm going to gift you with some totally unbiased considerations about bashML.
Pros:
- This is the Unix way (in fact, we just chained several programs from
coreutils) - Pipe'ing is efficient enough for a lot of use-cases
- Pipes don't require much memory
- Pipes define a directed graph (even though it's better to save intermediate results if you use them more than once)
- Scheduling is built-in
- Easy parallelization
- Commands are independent from one-another. When enough data is available to read from a pipe, they just run free.
parallelxargssort
Most importantly: no garbage collector.
Cons:
- Limited in what you can do (e.g.: you wouldn't make a neural network in bash)
- Limited in the kind of data you can handle
- Think quoted columns in a CSV file (unless you use some workaround like csvtool)
- No types
- Doing things in RAM is faster (if you are doing complex operations)
- Doesn't scale well to multiple machines
- Things can get pretty messy pretty fast
- Sad data scientists
Conclusion
There, you heard it here folks, you don't need Spark anymore. You hold the power of bash now. It has worked for humanity for 30+ years and it's here to stay. The next time you need to process some data ask yourself: why Spark when you can Bash?
On a more serious note, at rev.ng we actually ended up using this Unix-style approach because it is a sweet-spot for our use-case. Databases were way too slow and setting up some compute cluster seemed overkill for the amount of data we are working with, especially since we can actually do everything in bash on a single machine.
Now turn off you computer and go to sleep tell your friends about how bash people were doing machine learning before it was cool.
Cheers,